|
Many advances have been made in single-cell and single-molecule analysis. These technologies enable the detection of very small numbers of cells (e.g. infectious pathogens or cancers circulating in the bloodstream) or very small differences between cells (e.g. antibiotic or chemo resistance). But there remains a need for rapid and quantitative technologies that don't require up-front assumptions about which cells and molecules are of interest. For example, a patient with a bacterial bloodstream infection could harbor any combination of 200 common organisms. Moreover, every hour that the patient goes incorrectly diagnosed, their chances of survival goes down. The Fraley lab approaches this problem by developing techniques that require fewer assumptions and bias up front while still providing quantitative, sensitive, and specific details at the single cell level. We apply microfluidic, optical, engineering, and computational approaches to create enabling technologies that shed new light on complex host-pathogen interactions and outcomes.
|
Rapid Point-of-Care Diagnostics & Single Cell Analysis
|
Clinical Need: Neonatal Sepsis
|
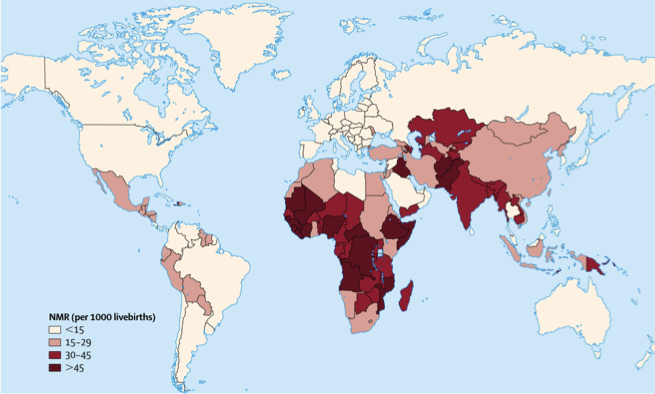
Variation between countries in neonatal mortality rates (NMR).
|
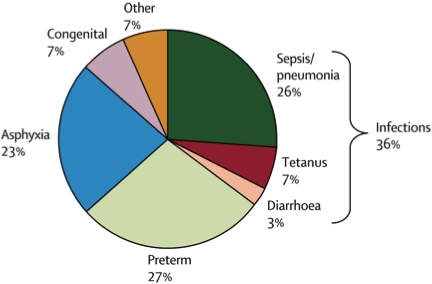
Estimated distribution of causes of neonatal deaths.
|
|
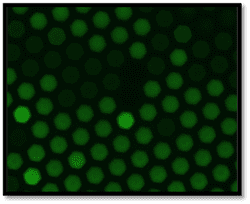
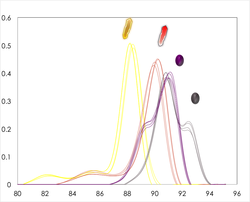
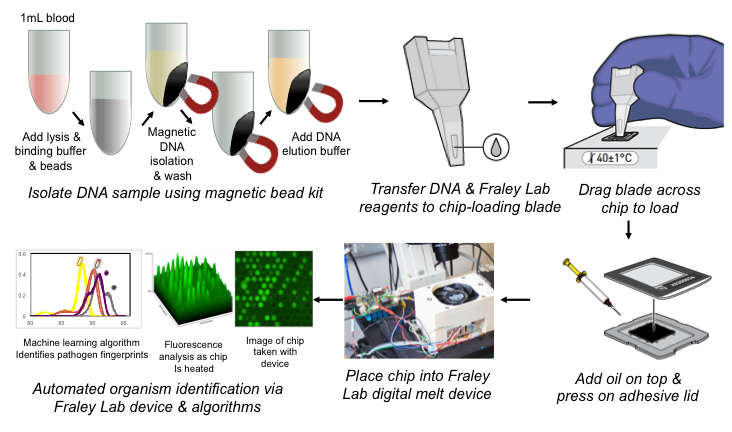
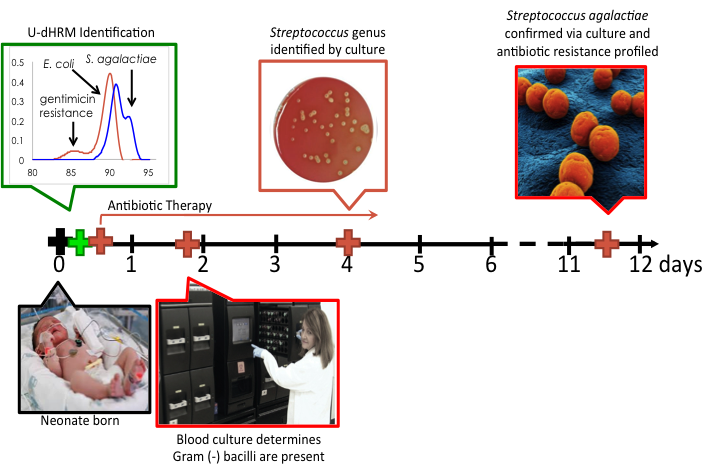
A typical clinical timeline of neonatal sepsis diagnosis and treatment is shown above. Our U-dHRM technology is poised to significantly improve time-to-diagnosis and enable accurate, targeted therapy.
|
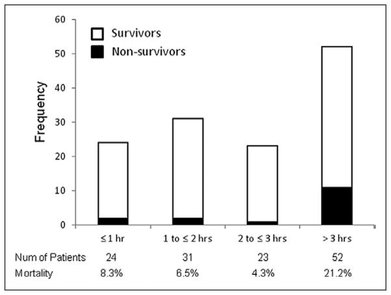
Time from symptom-based sepsis recognition to antimicrobial administration shows a 3 hour window for accurate treatment to be determined before mortality rate increases.
|
SI Fraley, et.al., Scientific Reports, 2015
P Athamanolap, et.al., PloS One, 2014
SI Fraley, et.al., Nucleic Acids Research, 2013
P Athamanolap, et.al., PloS One, 2014
SI Fraley, et.al., Nucleic Acids Research, 2013